Machine Learning Algorithms for the Identification of Cancer Cells Using Gene Expression Data
Dr Anders Jacobsen Skanderup, Adjunct Associate Professor, School of Computing
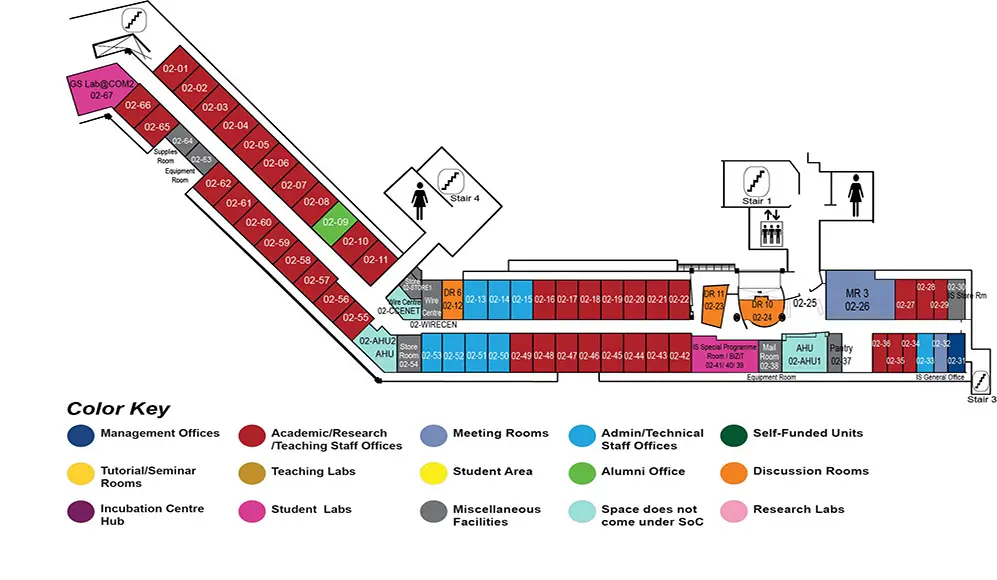
COM2 Level 2
MR3, COM2-02-26
ABSTRACT:
Cancer is one of the leading causes of death worldwide. Developing new computational techniques to understand cancer is one of the foremost research priorities. One of the actively researched subjects in cancer biology is the topic of tumor microenvironment - the study of the cellular composition of the tumor tissue and the interactions of the cells within it. In this thesis, we study and develop computational methods capable of providing high-level information about the cellular composition of the tumor microenvironment. Specifically, we develop a novel regression method for predicting tumor purity (proportion of cancer cells in a sample) from bulk gene expression data, benchmark it against existing methods, and extensively explore related validation strategies, and applications, as well as potential extensions of the approach.
On the highest level of complexity, tumors are heterogeneous masses composed of malignant (cancer) and non-malignant cells. Variation in tumor purity can both confound integrative analysis and enable studies of tumor heterogeneity. In this thesis, we develop and validate a machine learning-based method for accurate pan-cancer tumor PURity Estimation from gene Expression data - PUREE. PUREE, which uses a weakly supervised learning approach to infer tumor purity from a tumor gene expression profile was trained on gene expression data and genomic consensus purity estimates from approximately 8000 solid tumor samples and validated on several additional independent datasets. PUREE is able to predict purity with high accuracy across distinct solid tumor types and generalize to tumor samples from unseen tumor types and cohorts. Gene features of PUREE were further validated using single-cell RNA-seq data from distinct tumor types. In a comprehensive benchmark, PUREE outperformed existing transcriptome-based purity estimation approaches. Overall, we demonstrate that PUREE is a highly accurate and versatile method for estimating tumor purity and interrogating tumor heterogeneity from bulk tumor gene expression data.
Furthermore, we demonstrate how PUREE can be applied to microarray gene expression data, and how it can be modified further into a compact lab assay-based purity prediction method. We further explore how a method for identifying cancer cells from single-cell RNA-seq data can be constructed using machine learning techniques and how the feature set of PUREE can be used to potentially improve its performance.
In summary, in this thesis, we provide the necessary background for the topic of tumor microenvironment research and the computational techniques that are suitable for dissecting its complexity. We show how we created and validated a novel machine learning method to identify the proportions of cancer cells in tumor tissues using bulk gene expression data. We additionally show how to build machine learning models for identifying cancer cells using single-cell gene expression data, and how the feature set identified by the bulk gene expression-based method can be used in that task as well. We discuss the application of our approaches and provide an overview of future directions for similar work.

